
Moreover, just two classes of S2 binding antibodies have been described in the literature: those binding the fusion peptide and adjacent S2' cleavage site ( Dacon et al., 2022 Low et al., 2022 Sun et al., 2022) and those binding the S2 stem proximal to the viral membrane ( Hsieh et al., 2021 Pinto et al., 2021 Sauer et al., 2021 Zhou et al., 2022). Fewer than 5% of the ~7000 anti-SARS-CoV-2 spike monoclonal antibody sequences in the CoV-AbDab database bind S2 as of July 2022 ( Raybould et al., 2021). Whereas antibodies binding the spike RBD have been rigorously classified based on epitope recognized ( Barnes et al., 2020) and this information used to support development of RBD mosaic vaccines ( Cohen et al., 2022), a complementary analysis of the S2 domain is in its infancy. Moreover, the functionally analogous domains in the fusion proteins from influenza virus, respiratory syncytial virus, and human immunodeficiency virus are targeted by protective antibodies ( Impagliazzo et al., 2015 Corti et al., 2017), supporting the hypothesis that the spike S2 domain may also be an effective target. Early in the pandemic, S2-directed antibodies often dominated the immune repertoire in convalescent patients, indicating that at least some S2 epitopes are immunogenic ( Voss et al., 2021). In contrast to the S1 domain comprising the RBD and N-terminal domains, the S2 domain is highly conserved, with 63–98% sequence similarity in pairwise comparisons across the seven human coronaviruses ( Figure 1-figure supplement 1). Immunization with the entire spike ectodomain induces potently neutralizing antibodies that block binding of the receptor binding domain (RBD) to the ACE2 receptor ( Yuan et al., 2020), indicating that the RBD is a protective and immunogenic, as well as variable, antigen. The continued emergence of SARS-CoV-2 variants of concern underscores the need to identify therapeutic strategies more resistant to antigenic drift. Moreover, the seven coronaviruses known to infect humans are closely related to strains found in wildlife, foreshadowing future coronavirus outbreaks. This has culminated in the currently circulating Omicron subvariants with >30 amino acid changes that resist neutralization by all monoclonal antibodies with Emergency Use Authorization ( VanBlargan et al., 2022 Imai et al., 2023) and cause breakthrough infections in fully vaccinated individuals. Despite the successes of vaccines and antibody therapeutics that neutralize SARS-CoV-2 virus by disrupting interactions between the ACE2 receptor and the spike fusion protein, mutations accumulating primarily in the S1 domain have resulted in widespread evasion of antibodies elicited against early virus strains. The COVID-19 pandemic is the latest and largest of three deadly coronavirus outbreaks, including those caused by SARS-CoV in 2002 and MERS-CoV in 2012. 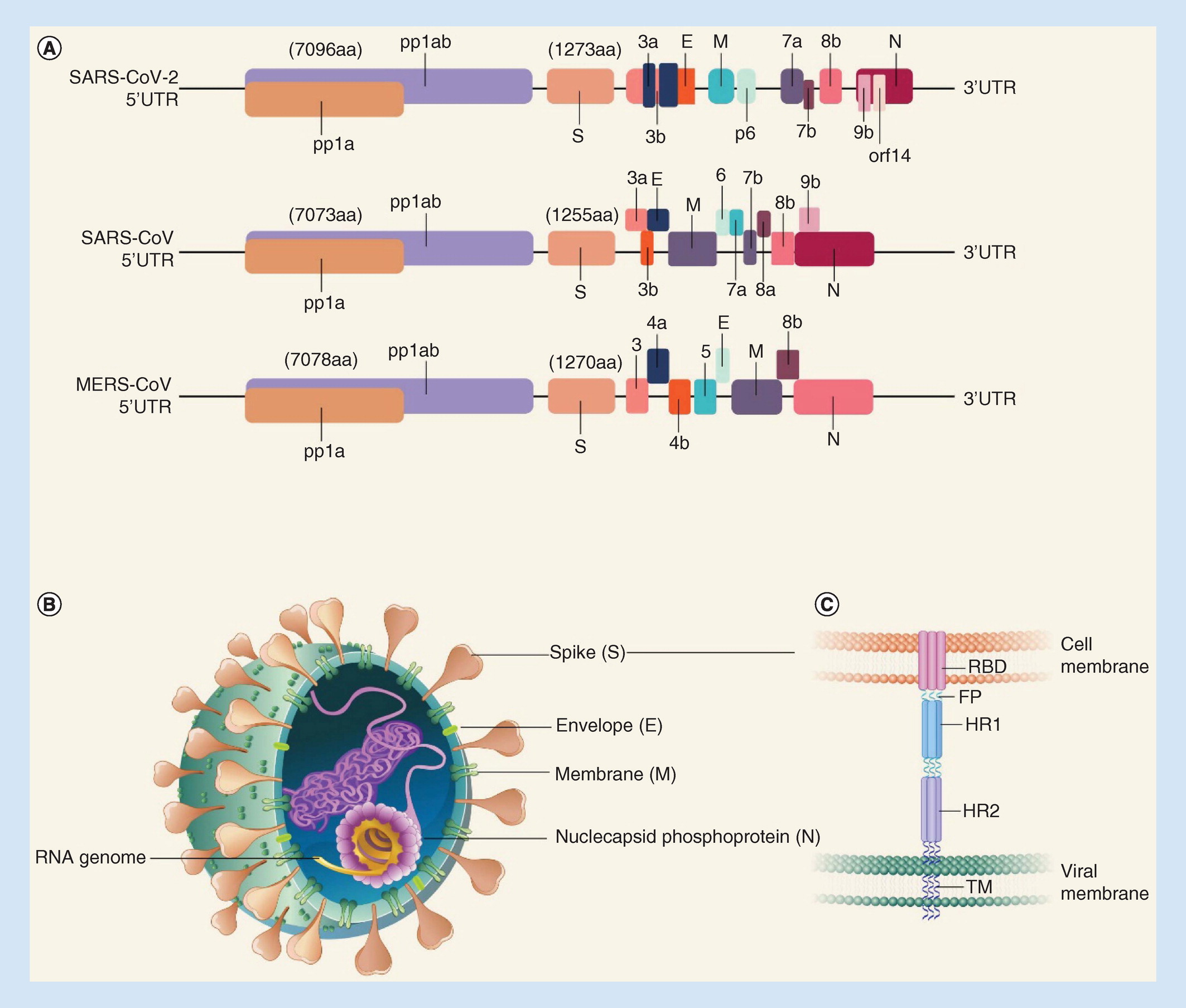
This work defines a third class of S2 antibody while providing insights into the potency and limitations of S2 core epitope targeting. Hinge epitope mutations that ablate antibody binding compromise pseudovirus infectivity, but changes elsewhere that affect spike opening dynamics, including those found in Omicron BA.1, occlude the epitope and may evade pre-existing serum antibodies targeting the S2 core. 
Antibody RAY53 binds the native hinge in MERS-CoV and SARS-CoV-2 spikes on the surface of mammalian cells and mediates antibody-dependent cellular phagocytosis and cytotoxicity against SARS-CoV-2 spike in vitro. We describe a highly conserved, conformational S2 domain epitope present only in the prefusion core of β-coronaviruses: SARS-CoV-2 S2 apex residues 980–1006 in the flexible hinge. To address the ongoing SARS-CoV-2 pandemic and prepare for future coronavirus outbreaks, understanding the protective potential of epitopes conserved across SARS-CoV-2 variants and coronavirus lineages is essential.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
